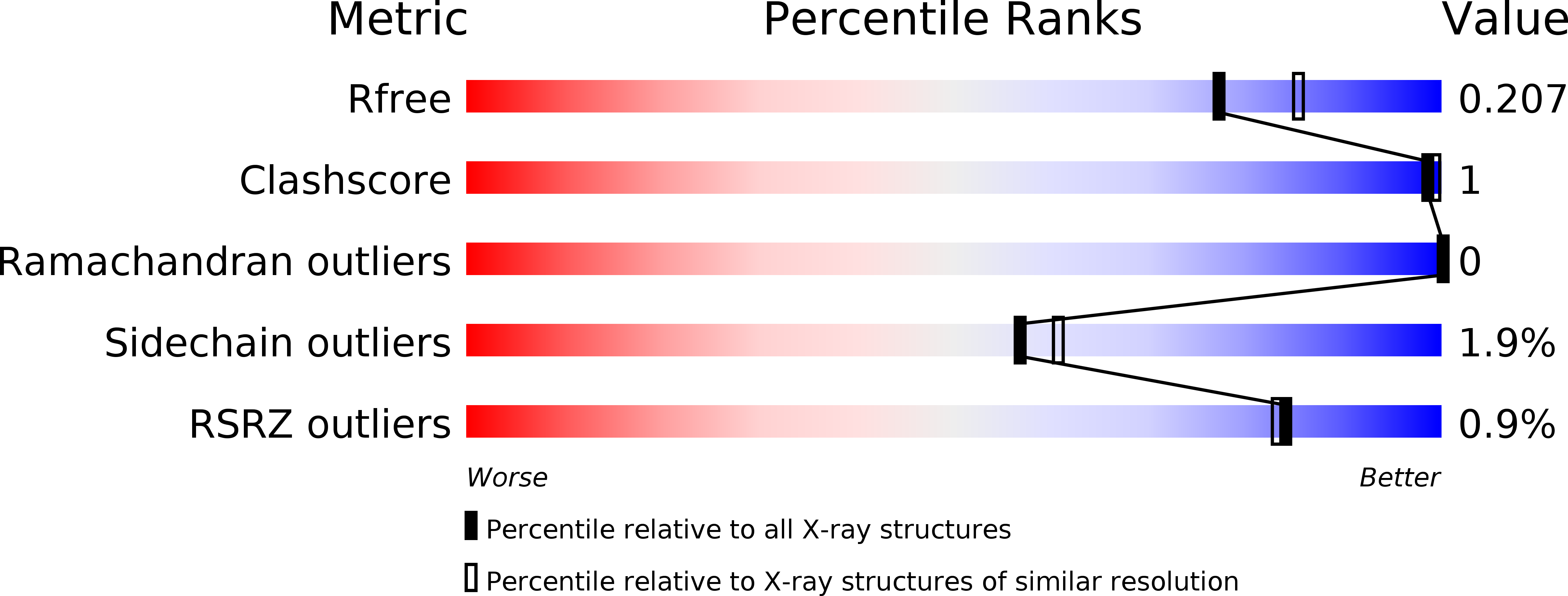
Discovery of Inhibitors of Four Bromodomains by Fragment-Anchored Ligand Docking.
Marchand, J.R., Dalle Vedove, A., Lolli, G., Caflisch, A.(2017) J Chem Inf Model 57: 2584-2597
- PubMed: 28862840
- DOI: https://doi.org/10.1021/acs.jcim.7b00336
- Primary Citation of Related Structures:
5OR8, 5OR9, 5ORB - PubMed Abstract:
The high-throughput docking protocol called ALTA-VS (anchor-based library tailoring approach for virtual screening) was developed in 2005 for the efficient in silico screening of large libraries of compounds by preselection of only those molecules that have optimal fragments (anchors) for the protein target. Here we present an updated version of ALTA-VS with a broader range of potential applications. The evaluation of binding energy makes use of a classical force field with implicit solvent in the continuum dielectric approximation. In about 2 days per protein target on a 96-core compute cluster (equipped with Xeon E3-1280 quad core processors at 2.5 GHz), the screening of a library of nearly 77 000 diverse molecules with the updated ALTA-VS protocol has resulted in the identification of 19, 3, 3, and 2 μM inhibitors of the human bromodomains ATAD2, BAZ2B, BRD4(1), and CREBBP, respectively. The success ratio (i.e., number of actives in a competition binding assay in vitro divided by the number of compounds tested) ranges from 8% to 13% in dose-response measurements. The poses predicted by fragment-based docking for the three ligands of the BAZ2B bromodomain were confirmed by protein X-ray crystallography.
Organizational Affiliation:
Department of Biochemistry, University of Zürich , CH-8057, Zürich, Switzerland.